Plant Genetics Meets Gut Microbiome: How Crop Diversity May Unlock Novel Prebiotics
By Qinnan Yang, PhD
Prebiotics are getting great attention for good reason—they can help steer our gut microbes and metabolism in ways that can boost immunity, keep our body running smoothly, and even prevent certain diseases. However, as the prebiotic world draws increased interest, new questions are popping up: how do we identify novel prebiotics, and can we produce them in a sustainable and relatively accessible way?
The breeding target of modern crops is largely still about yield, whether directly by making high-yield crops or indirectly by improving resilience to harsh environments, pests, or diseases. This leaves a huge knowledge gap in understanding the health-related traits in crops. With this huge gap comes mounting opportunity: a systematic approach to catalogue and identify these health-promoting traits that lie under the vast amount of genetic resources is often overlooked. We chose to study sorghum, an ancient grain that uses fewer natural resources in its production and is one of the model sustainable crops. Sorghum is high in dietary fiber and antioxidants and carries an enormous amount of genetic and phenotypic variation. These characteristics make this overlooked grain perfect for identifying novel prebiotics.
Our research group at Nebraska Food for Health Center uses the classic forward genetics approach to find genetic variation in food crops that affect the human gut microbiome and further find the genes and potentially bioactive components (prebiotics) in sorghum. The process of this forward genetics has three major components: 1) traits of the crop that can be accurately measured; 2) a crop population with rich diversity on the trait of interest; 3) an advanced mapping algorithm that can identify the genes that affect the traits.
Effects on the gut microbiome as health traits of sorghum
You are what you eat, and microbes that reside in your gut respond even more dramatically to a bowl of oatmeal versus a slice of pizza. The gut microbiome can be associated with numerous metabolic and inflammatory diseases, thus the effect of a specific food crop on the microbiome can naturally serve as health-related traits. In a perfect world, the gut microbiome affecting traits of a food crop can be measured by conducting trials on participants eating this food crop. Unfortunately, it is very challenging and almost impossible to catalogue and identify plant genetic variations that affect the gut microbiome, which is simply due to the large number of crop varieties that need to be tested.
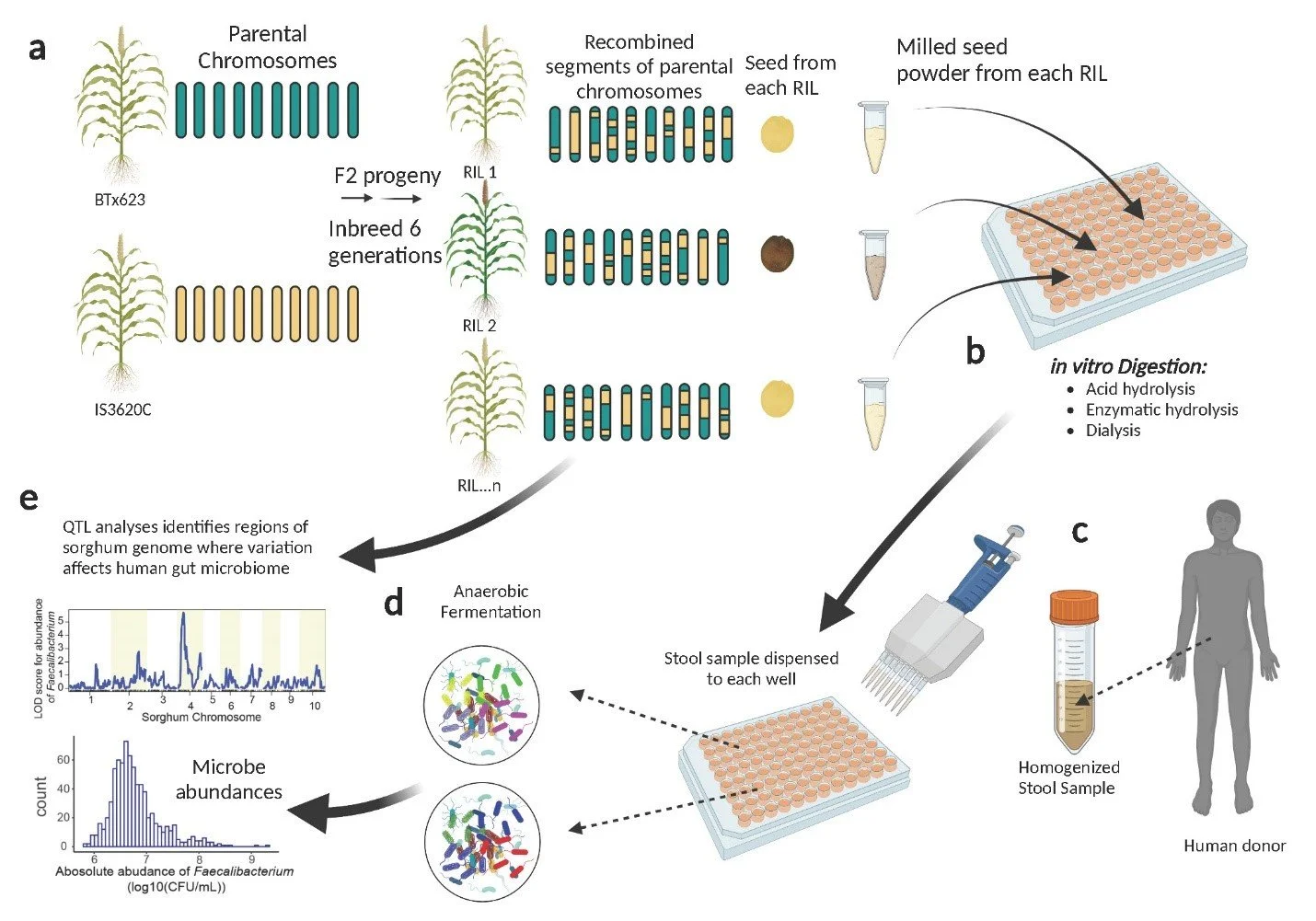
Instead, we developed a state-of-the-art automated in vitro microbiome screening (AiMS) to measure the interaction of human gut microbes with pre-digested grain (Figure 1). The AiMS method takes grain samples from each sorghum line and puts them through a process that’s a lot like what happens in our digestive system. First, the grains are milled and then treated with acids and digestive enzymes to simulate digestion, and finally, dialysis is used to mimic absorption in the small intestine. After this mock digestion, what’s left of the grain is combined with samples of gut microbes (taken from a human donor’s stool) and incubated in an oxygen-free environment, recreating conditions in the colon. Lastly, we use 16S rDNA sequencing to measure which microbes thrive in each fermentation setup. These microbial profiles become data points—quantitative traits.
Figure 1. Diagram of the AiMS for genetic analysis of microbiome traits. Adatped from [1] Yang, Q., Van Haute, M., Korth, N., Sattler, S. E., Toy, J., Rose, D. J., ... & Benson, A. K. (2022). Genetic analysis of seed traits in Sorghum bicolor that affect the human gut microbiome. nature communications, 13(1), 5641
Figure 2. Sorghum grew in the greenhouse at the University of Nebraska-Lincoln with futuristic purple light.
Collection of sorghum lines with distinct phenotypes
For this proof-of-concept study, we chose to use a group of well-documented 294 sorghum recombinant inbred lines (RILs), essentially, sorghum lines that were bred from two genetically different parents. This population of lines shows a great difference in agronomic traits such as plant height and flowering time. The color of sorghum grain from this population is also very different.
As we all know, the environment could have a huge impact on crops. I grew this sorghum population in a greenhouse with controlled temperature, humidity and even daytime (photoperiod). We took grain from each RIL and used our automated in vitro microbiome screening (AiMS) technique to see how human gut microbes interact with the pre-digested grain.
Quantitative trait Locus (QTL) Mapping identifies sorghum genome regions that affect gut microbiome
The mapping was done by matching differences in the sorghum genome to changes in gut microbe abundance. Changes in short chain fatty acids, which play vital roles in maintaining the intestinal barrier and reducing inflammation, were also analyzed. This analysis revealed which genetic regions in sorghum influence the way its digested grain affects the gut microbiome. We found 26 genetic regions—called quantitative trait loci, or QTLs—that were significantly linked to changes in the abundance of 19 different types of gut microbes. These QTLs were spread across nine out of ten chromosomes, showing that many genes collectively help shape how sorghum grain influences the human gut microbiome.
Condensed tannin is regulated by two QTLs and is identified as potential prebiotic to increase Faecalibacterium
Two genetics loci that are on chromosomes 2 and 4 influenced 8 different bacteria genera, including Faecalibacterium (a butyrate producing bacteria and has anti-inflammatory properties). The same two loci were also found to control condensed tannin, also known as proanthocyanidin, a type of polyphenol that shows great antioxidant activity), making seed color darker.
We therefore hypothesize that condensed tannin increased Faecalibacterium. To support our hypothesis, we did a series of experiments that showed: 1) Three high condensed tannin sorghum lines increased the abundance of Faecalibacterium compared to their corresponding near-isogenic parental lines; 2) Complementing condensed tannin to the non-tannin sorghum expands the Faecalibacterium abundance; 3) condensed tannin alone without sorghum can support Faecalibacterium growth.
A practical pipeline that can be used in other food crops to identify novel plant-based prebiotic
Our study shows how this novel methodology can pinpoint specific genes in food crops that influence both seed composition and how those seeds interact with the human gut microbiome. The great news? This method could work for any food crops, helping us build a detailed map of seed traits that shape our gut microbes. We’re optimistic that, in the future, crop breeders will be able to focus on traits that boost beneficial gut bacteria—opening doors to new ways crops can support human health.
References
1. Yang, Q., Van Haute, M., Korth, N., Sattler, S. E., Toy, J., Rose, D. J., ... & Benson, A. K. (2022). Genetic analysis of seed traits in Sorghum bicolor that affect the human gut microbiome. nature communications, 13(1), 5641
2. Korth, N., Yang, Q., Van Haute, M. J., Tross, M. C., Peng, B., Shrestha, N., ... & Benson, A. K. (2024). Genomic co-localization of variation affecting agronomic and human gut microbiome traits in a meta-analysis of diverse sorghum. G3: Genes, Genomes, Genetics, 14(9), jkae145.
3. Suhr Van Haute, M. J., Yang, Q., Korth, N., Happ, M. M., Kok, C. R., Miao, C., ... & Benson, A. K. (2025). Genetic variation and historical breeding patterns in common bean (Phaseolus vulgaris L.) affect fermentation patterns by the human gut microbiome. Communications Biology, 8(1), 1690.